
A new article co-authored by Mattias Jakobsson and Max Larena has been published in Immunogenetics.
Schaschl, H., Herzog, T., Oberreiter, V. et al. HLA diversity and signatures of selection in the Maniq, a nomadic hunter-gatherer population in Southern Thailand. Immunogenetics 77, 23 (2025). https://doi.org/10.1007/s00251-025-01380-0
Abstract
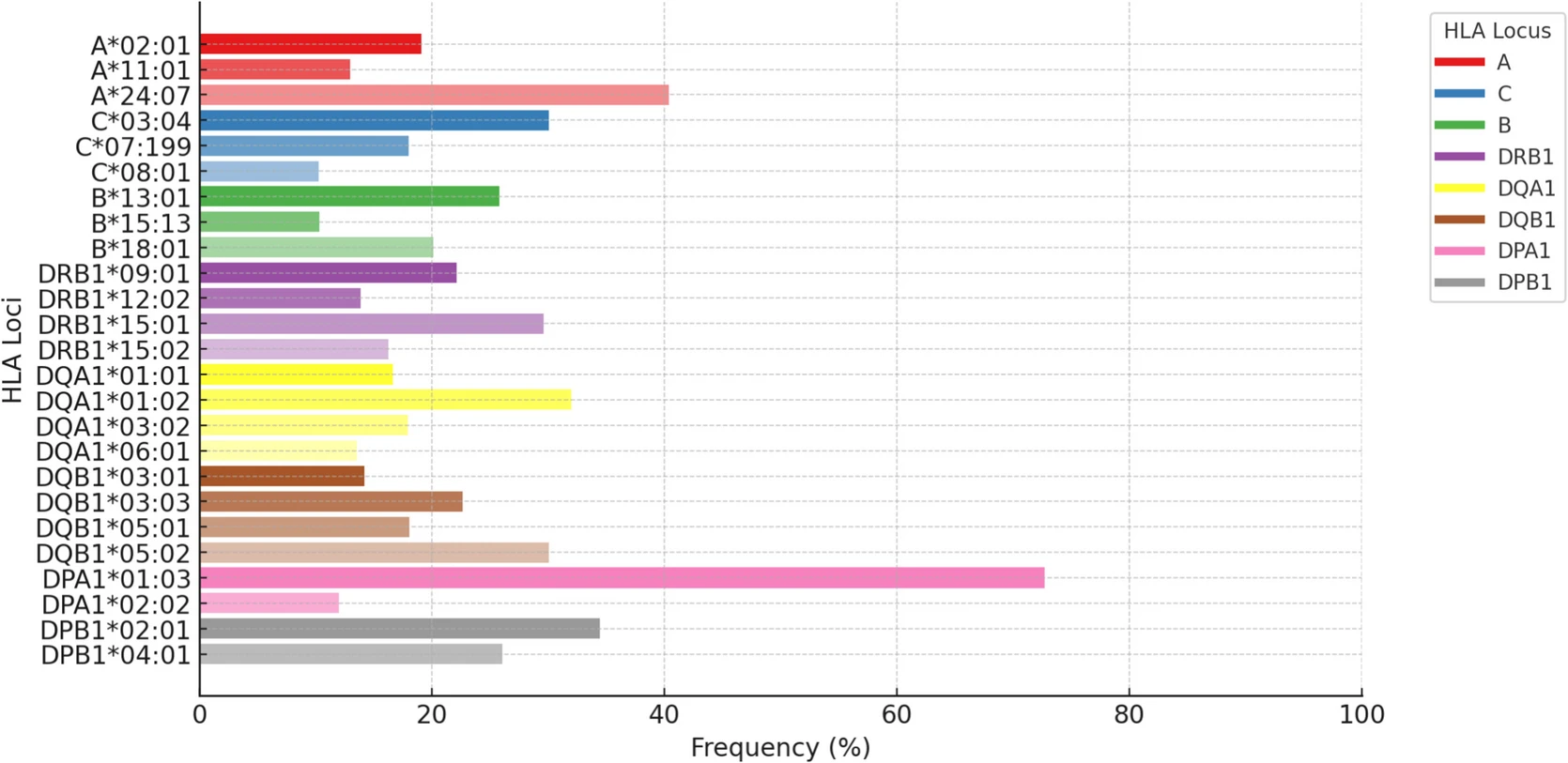
The Maniq, a small and isolated nomadic hunter-gatherer population from the rainforests of Southern Thailand, offer a unique context for investigating how demographic history, genetic drift, and pathogen-driven selection shape human leucocyte antigen (HLA) diversity. Using high-coverage whole-genome data from 21 individuals (12 unrelated), we genotyped HLA alleles with HLA-HD and T1K, identifying 32 alleles in classical and 14 in non-classical HLA genes. Although overall HLA diversity was comparatively low, a few alleles at each locus occurred at high frequency, mirroring patterns observed in other small, isolated populations. Principal-component analysis clustered the Maniq with other Austroasiatic-speaking Semang hunter-gatherers (Jehai, Kintaq) on the Malay Peninsula and, intriguingly, with the Austronesian-speaking Tao of Taiwan, indicating shared immunogenetic features across linguistic boundaries. Despite reduced diversity, multiple loci bore signatures of both long-term balancing and recent positive selection. Several SNPs under selection were in complete linkage disequilibrium with eQTLs known to influence responses to hepatitis B virus (HBV) and other pathogens, suggesting that pathogen-driven pressure—in particular HBV—may have contributed to recent HLA evolution in the Maniq. These findings provide critical insights into how demographic constraints and pathogen landscapes converge to shape HLA diversity and evolution. In light of increasing infectious disease burdens in indigenous communities, our results underscore the importance of studying small, isolated populations to better understand the adaptive significance of HLA genes.

